7. Minding your Rs and Qs I
Written by Norm MacLeod - The Natural History Museum, London, UK (email: n.macleod@nhm.ac.uk). This article first appeared in the Nº 61 edition of Palaeontology Newsletter.
Part 1
All the methods we've been discussing to this point in the PalaeoMath 101 essays have exhibited an underlying similarity. No, it's not only the fact that all have made extensive use of the four canonical arithmetic operators (+, -, x, /). Rather, our discussions of bivariate regression, multiple regression, principal component analysis (PCA), and factor analysis (FA) are united by the fact that each technique has focused its mathematical sights on relations between variables, the sets of observations or measurements we make on specimens, at localities, etc. In particular, all these methods have operated on values, or matrices of values, of the covariance or correlation coefficients.
Recall the definitions of these quantities. The variance (first discussed in the Regression 2 essay from Newsletter 56) is the sum of the squared deviations of the measurements from their mean, divided by one less than the number of measurements in the sample. The covariance (Newsletter 56) is the joint variation of two variables around their common mean, and the correlation coefficient (Newsletter 58) is the ratio of two variable's covariance to the product of their standard deviations. All these quantities can be used to make statements about the character of the distribution of measurements in populations, provided the samples used in the calculation of these quantities provide accurate representations of the underlying population of possible measurements. Thus, the structure of variance-covariance relations among the three variables of our trilobite dataset...
| Genus | Body Length (mm) | Glabellar Length (mm) | Glabellar Width (mm) |
|---|---|---|---|
| Acaste | 23.14 | 3.50 | 3.77 |
| Balizoma | 14.32 | 3.97 | 4.08 |
| Calymene | 51.69 | 10.91 | 10.72 |
| Ceraurus | 21.15 | 4.90 | 4.69 |
| Cheirurus | 31.74 | 9.33 | 12.11 |
| Cybantyx | 26.81 | 11.35 | 10.10 |
| Cybeloides | 25.13 | 6.39 | 6.81 |
| Dalmanites | 32.93 | 8.46 | 6.08 |
| Delphion | 21.81 | 6.92 | 9.01 |
| Ormathops | 13.88 | 5.03 | 4.34 |
| Phacopdina | 21.43 | 7.03 | 6.79 |
| Phacops | 27.23 | 5.30 | 8.19 |
| Placopoaria | 38.15 | 9.40 | 8.71 |
| Pricyclopyge | 40.11 | 14.98 | 12.98 |
| Ptychoparia | 62.17 | 12.25 | 8.71 |
| Rhenops | 55.94 | 19.00 | 13.10 |
| Sphaerexochus | 23.31 | 3,84 | 4.60 |
| Toxochasmops | 46.12 | 8.15 | 11.42 |
| Trimerus | 89.43 | 23.18 | 21.52 |
| Zacanthoides | 47.89 | 13.56 | 11.78 |
| Mean | 36.22 | 9.37 | 8.98 |
| Variance | 346.89 | 27.33 | 18.27 |
... can be represented by the covariance and/or the correlation matrices of those measurements...
| Covariance | Correlation | |||||
|---|---|---|---|---|---|---|
| Body Length | Glabella Length | Glabella Width | Body Length | Glabella Length | Body Length | |
| Body Length | 329.829 | 87.191 | 64.416 | 1.000 | 0.895 | 0.859 |
| Glabella Length | 87.191 | 27.333 | 20.315 | 0.895 | 1.000 | 0.909 |
| Glabella Width | 68.416 | 20.315 | 18.266 | 0.859 | 0.909 | 1.000 |
...and the structure of these relations' including representations of between-specimen relations' can be summarized numerically and even portrayed graphically in the form of scatterplots along either variable or composite variable axes.
There is another way of looking at the problem of summarizing structural relations within this dataset, however. Suppose we're not really interested in the structure of the variables' relations with one another, but instead wanted to know about the structure of similarity relations between individual trilobite specimens. Of course, this is not an idle consideration. In many instances relations among objects"or populations that can be represented by a specimen or locality"are precisely what we do want know about. From our previous discussions you will recall that placement of the PCA and FA axes we used to portray inter-specimen relations was determined entirely on the basis of the structure of the covariance or correlation matrices. The tables of eigenvalues we generated as part of those operations tell us what proportion of the observed variance is being represented along the various PCA and FA axes. That's all more-or-less straight-forward. But how "good" were those pictures of inter-specimen similarities and differences that were presented on those plots of specimen points projected onto the PCA and FA axes? Those PCA and FA axes reproduce the structure of the covariance or correlation matrices. They do not sense, much less optimize, inter-specimen relations per se. Should we always believe the picture they paint of inter-specimen relations? How can we determine whether the analysis introduced sufficient distortion to bias any interpretation we may wish to make? Are we always restricted to using the covariance or correlation matrix as a proxy for measuring inter-object similarity?
Interestingly, a realization of what PCA and FA actually do implies that there exists a 'parallel universe' of methods, similar in form to PCA and FA, but different in that their focus would be directed to summarizing inter-specimen relations. Such a parallel universe does indeed exist within numerical data analysis, and with it tantalizing questions about the particular strengths of the methods that populate that universe and the character of the mathematical interface between those two universes. It is this alternative universe of specimen-based (as opposed to variable-based) methods we'll explore in the next few essays.
As with all things scientific, this new universe must be given a name. When we are basing a multivariate procedure on the structure of relations among variables, data analysts say they are working in the R-mode. Up to now we have been concerned exclusively with R-mode data-analysis techniques. With this essay I'l introduce some of the methods data analysts use to undertake investigations based on the structure of relations among specimens or localities. This is work undertaken in the 'Q-mode'. Where did these terms come from? They are mathematical symbols for the two different types of structure matrices. The 'R' in R-mode refers to the R that linear algebrasts use to represent the correlation matrix in equations. At least this makes some sense insofar as rij is the standard symbol for the correlation between variables i and j. Why 'Q' for the matrix of relations among objects (across variables)? As it turns out Q is the symbol mathematicians use to represent a distance or association matrix. Most R-mode techniques have Q-mode counterparts, though in this essay we'll see that sometimes this apparent correspondence is more a matter of rhetoric than reality. In any event, I'll focus on the Q-mode counterparts to PCA and FA in this essay. Once we're comfortable with these we'll go on to consider even more interesting techniques that provide properly trained investigators with the power to perform R-mode and Q-mode analyses simultaneously.
The first thing"some would say the main thing"to understand about Q-mode analyses is that there are a far larger selection of similarity indices available for assessing the structure of relations between objects than for assessing the structure of relations between variables. Literally hundreds of Q-mode similarity indices have been proposed. Discussing in detail even those one sees referred to repeatedly would be an essay in itself and reviewing the entire literature would require a particularly tedious book. In this essay we'll focus on just three of these Q-mode indices to give a flavour of the available range.
Before we do that, though, it might be good to consider the question "Why not just regard the specimens as variables, the variables as specimens"in other words, transpose the data matrix"and use the normal covariance or correlation coefficients as the basis for a normal PCA or FA analysis?†After all, those methods are well established, the characteristics of these indices well understood, and, so long as one is willing to make the concept transpositions mentioned above, the calculations are possible. What's the problem with calculating the covariance between objects?
Most authors of data analysis textbooks either don't consider this question at all or tend to dismiss it with cryptic comments like "The reasoning behind such a measure is at best obscure." (Davis 2002, p. 540). The actual reasons are simple to understand but may not be as obvious to neophytes as some experienced practitioners might believe. The covariance coefficient is derived from the variance coefficient which is itself a statement about the character of a group of observations. The sample variance of glabellar length is a proxy summary of all observations of this quantity that could have been made available to the observer. In calculating the variance one presumes the units of measurement have are the same (or have been made the same) for each observation. One could calculate the 'variance' of such a set of observations, some of which had been made in centimetres, some in millimetres, some in inches, and some in feet. The result, though, would indicate more about differences in the units of measure within the dataset than about any biological reality. While it is relatively easy to control for this obvious source of inconsistency within the context of a single variable, it is much more difficult, sometimes impossible, to undertaken such standardizations for a group of intrinsically different variables. In such instances selection of an index that takes differences between variables into consideration in known and logical ways is necessary. Another reason why a special class of Q-mode similarity indices are necessary is that a single set of observations from a population can be expected to be drawn from a frequency distribution of known, or at least ascertainable, shape. The variance index of a sample can be related directly to the shape of a distribution of observations in the sample's parent population (see the Regression 3 essay from Newsletter 59). A 'variance' calculated from a mixture of samples from different distributions, often without scale or unit in common, will obey Chebychev's theorem, but not its corollary for more normal samples. The final reason is the simple, practical expedient that in some cases one may want to use a similarity index that differentially weights some aspect of the data known a priori to be relevant to the problem at hand. For all these reasons the blind use of covariance or correlation indices as the structural basis for a Q-mode analysis is ill-advised. Unfortunately, both public-domain and commercially available data analysis packages cannot prevent the use of inappropriate similarity indices. Since the responsibility for selecting the correct method always lies with the data analyst, practitioners (and reviewer's of practitioner's work) should always state the nature of the basis matrix and the reasons they chose the similarity index used to determine that matrix.
So, what are some of the more typical Glabella Width-mode similarity indices and when should they be used? Perhaps the most straightforward Q-mode index is the squared Euclidean Distance.
(7.1)
dij=p∑k=1(xik−xjk)2
In this equation i represents the ith specimen, j represents the jth specimen, and p represents the total number of variables measured on each specimen. Geometrically, this quantity represents the simple straight-line distance between two specimens in multivariate space. Using equation 7.1 to assess the structure of distances between the first three taxa listed in Table 1 yields the following matrix.
| Acaste | Balizoma | Calymene | |
|---|---|---|---|
| Acaste | 0.000 | 78.109 | 918.313 |
| Balizoma | 78.109 | 0.000 | 1488.770 |
| Calymene | 918.313 | 1488.770 | 0.000 |
Note the difference in the structure of this matrix from those we've dealt with previously. The trace of the matrix is filled with zeros because the distance of any object from itself is 0.0. Large distance coefficients (e.g., the distance between Balizoma and Calymeme) indicate specimens that are very different from one another, while smaller coefficients (e.g., the distance between Acaste and Balizoma) indicate specimens that are comparatively similar, at least in terms of the measurements taken or observations made. This is very different from the covariance and correlation matrices where large values meant high degrees of similarity and small values meant substantial difference. Distance matrices are often referred to as 'dissimilarity' matrices because the magnitudes of the values express relative degrees of difference rather than similarity. This distinction having been made, distance matrices, like similarity matrices, are all symmetrical about their trace.
The squared Euclidean Distance is perhaps the most commonly used Q-mode distance index and is used primarily for data composed of continuous variables of the same type (e.g., distances between landmarks) and measured in the same units (e.g., mm). Still, many data matrices one might want to analyze are composed of different types of variables, often measured in different and incompatible units. In R-mode analyses we handle this situation by using the correlation coefficient because this index standardizes the data for differences in magnitude (and hence the variance) of the measurements. Standardization is often desirable in Q-mode analyses, but the mathematical operations are more complex and can be applied at different points in the analysis.
At the level of the similarity index, if we want to stick with a distance-based similarity measure we need to apply a scaling correction to the different variables during calculation of the component differences between all pairs of specimens. The most popular of the scale-corrected distance indices is the Gower Coefficient.
(7.2)
sij=p∑k=11−|xik−xjk|/Rkp
In this expression Rk represents the range of the kth variable. The Gower index is based on a distance index (|xik−xjk|/Rk) but subtracting this distance from 1.0 forces its range to fall within an interval from 0.0 to 1.0 with 1.0 representing perfect identity. Q-mode matrices in which the trace is occupied by 1s are called association matrices. Although the Gower matrix may look like a correlation matrix, the calculations are very different and, unlike the correlation coefficient, there is no mirroring of the index across the scale origin to represent inverse similarity (though the Gower Coefficient can be adjusted easily to incorporate this feature). Table 3 illustrates values of the Gower matrix for the first three taxa in Table 1.
| Acaste | Balizoma | Calymene | |
|---|---|---|---|
| Acaste | 1.000 | 0.947 | 0.618 |
| Balizoma | 0.947 | 1.000 | 0.593 |
| Calymene | 0.618 | 0.593 | 1.000 |
An interesting and useful feature of the Gower index is that it can be used not only with incompatible continuous variables (say, distances and volumes), but with sets of variables of any type (e.g., continuous measured variables, ratios, integer counts, nominal variables) with the only constraint being that all variables must be represented in numerical form. Of course, the manner of coding and relevance of all variables always needs to be justified and inter-variable differences always need to be kept in mind when interpreting a Gower matrix or summaries of its structure (see below). Nevertheless, the ability to include diverse sets of variables in the same analysis should the need arise is a powerful and attractive feature of Gower matrix-based Q-mode analyses.
Although the Gower index approaches the concept of the R-mode correlation coefficient in terms of imposing limits on the index's range, it retains a degree of sensitivity to the magnitude of the observations. For example, analyses of a set of fossils that were very similar in shape, but differed in size, would produce a Gower matrix that was structured according to those size differences. A different Q-mode similarity index is needed if we are to eliminate differences in the magnitude of constituent variables from consideration entirely. The most popular index that performs this task, called the Cosine θ, or 'Cosine theta', index, employs a geometric concept identical to that of the correlation coefficient.
The Cosine θ index is calculated as follows.
(7.3)
cosθij=p∑k=1xikxjk√p∑k=1x2ikx2jk
The Cosine theta index equation is actually a generalized expression for determining the angle between two vectors in multivariate space. As such, it focuses its assessment of similarity entirely on the directionality of the vectors and ignores differences in their length. It is also numerically equivalent to the Pearson product-moment correlation coefficient if there are only two variables and if the data are mean centered and standardized against the standard deviation.
The effects of this partitioning of the similarity into magnitude-related and non-magnitude-related parts can be striking. Table 4 illustrates values of the Cosine θ index for the first three taxa in Table 1.
| Acaste | Balizoma | Calymene | |
|---|---|---|---|
| Acaste | 1.000 | 0.987 | 0.998 |
| Balizoma | 0.987 | 1.000 | 0.996 |
| Calymene | 0.998 | 0.996 | 1.000 |
First, note that the Cosine θ index, like the Gower index, does not express dissimilarity (as all distance indices do), but true similarity with values close to the upper limit of the index (1.0) representing data patterns that are almost identical to one another. The trace of the matrix is filled with 1s denoting the perfect identity of any specimen with itself. The Cosine theta index also allows inverse similarity to be represented insofar as its natural bounding interval stretches from -1.0 to +1.0. Perhaps even more surprising, though, is the manner in which the Cosine θ index has represented similarity relations in our trilobite data. In both the Euclidean and Gower matrices (Tables 2 and 3, respectively) the most similar specimens were identified as Acaste and Balizoma while the least similar identified as Balizoma and Calymeme. The Cosine θ matrix shows similarity relations among these taxa to be reversed. Why is this? Inspection of Table 1 reveals the answer. The absolute magnitude of the three measurements for Calymeme is more than twice the magnitudes of the corresponding measurements for Acaste and Balizoma. Thus, if the scale of the measurements is considered important Acaste and Balizoma are more similar to each other than either is to Calymeme. But if scaling differences are deemed to be nuisance factors and differences in the magnitude of the measurements eliminated from consideration, Acaste is more similar to Calymeme than to Balizoma. This reversal of fortunes, so to speak, illustrates the power that comes from being able to 'fine-tune' the representation of similarity that is characteristic of Q-mode analysis. It also shows how important it is to get the similarity index right in designing a Q-mode data analysis. Alternatively, if one simply wants to probe the role of scaling factors in controlling patterns of overall similarity, a Cosine θ approach, in conjunction with a distance-based approach, would be indicated.
Once you've decided which Q-mode similarity index is appropriate for your data, the steps necessary to summarize the structure of this matrix, achieve dimensionality reduction, and produce images of the dominant similarity relations should begin to look familiar. Say, for example, you wanted to perform an analysis that focused on reproducing the matrix of similarities among specimens, portray that structure in a space of reduced dimensionality, and ensure the variables comprising that space were uncorrelated with one another. This would imply use of the Q-mode analogue to principal components analysis (PCA), which is termed Principal Coordinates Analysis (PCoord). Since all the trilobite measurements are distances I'll choose to preserve scaling distinctions among taxa by basing my analysis on the squared Euclidean Distance matrix.
One disadvantage of Q-mode analyses is that, because one usually collects data from more specimens than variables, specimen-based similarity matrices are larger than variable-based matrices and thus entail more computations. This was a much more serious problem in the past, when computer memories were far smaller than they are today, but one still sees allusions to the 'inefficiency' of Q-mode methods in published discussions. One limitation that is still with us, however, is the space required to show various data matrices. Accordingly, please see the PalaeoMath 101 Spreadsheet for a full listing of the squared Euclidean Distance matrix for all 20 trilobite species.
Once the squared Euclidean Distance matrix for the dataset has been obtained it is usually standardized according to the following equation.
(7.4)
aij=¯di+¯dj−¯¯d−dij2
This ensures the origin of the resulting PCoord axis system will coincide with the centroid of the data's point cloud in multidimensional space and 'closes' the similarity matrix by forcing the values of all rows and columns to sum to 0.0. The transformed similarity matrix is then decomposed using eigenanalysis in a manner identical to that used for PCA.
The eigenvalues obtained from the trilobite dataset using this method are listed below.
| Principal Coord. | Eigenvalue | Percent | Cum. Percent |
|---|---|---|---|
| 1 | 7278.440 | 97.601 | 97.601 |
| 2 | 141.677 | 1.900 | 99.501 |
| 3 | 37.217 | 0.499 | 100.00 |
There are several things to note about this matrix. First, although the Euclidian Distance matrix has a dimensionality of 20 rows and 20 columns, there are only three positive eigenvalues. These mirror the three eigenvalues obtained from the R-mode covariance matrix for these data (see PalaeoMath 101, Principal Components Analysis (Eigenanalysis & Regression 5), Table 5, Newsletter 59). The smaller-than-expected number of positive eigenvalues is due to the fact that, despite the number of taxa included in the dataset, only three measurements were taken. It is generally the case that the eigenanalytic decomposition of a Q-mode distance or association matrix will yield only as many positive eigenvalues as variables actually measured2. Second, the magnitude of the eigenvalues themselves are much larger than the corresponding eigenvalues for the R-mode PCA analysis of these data. This also reflects the difference in the nature of the similarity matrices subjected to eigenanalysis. In the case of the R-mode analysis of these data the covariance values ranged from 18.27 to 346.89 whereas the corresponding figures for the standardized distance matrix are -1,306.79 and 3,179.42. Third, despite these clear and easily appreciated differences, the percent contribution of each principal coordinate to characterizing the squared Euclidean Distance matrix is precisely the same as the percent contribution of each principal component to the characterization of the covariance matrix for these data. This identity conforms to Gower's (1966) proof that a PCoord of a Q-mode squared Euclidean Distance matrix is an exact mirror, or 'dual', of the covariance-based PCA for the same data. In passing, it is also worth mentioning that by moving to an analysis of relations between specimens as opposed to variables we are no longer using eigenanalysis to summarize variance, but multivariate distance. In a sense this is simply a rhetorical distinction. The calculations use for summarizing the contribution of each principal component-coordinate to the basis matrix is identical in PCA and PCoord. Nevertheless, the nature of the similarity matrix always needs to be kept in mind.
Since there are only three positive eigenvalues that can be extracted from the matrix of squared Euclidean distances between trilobite taxa, we only need to consider the eigenvectors that correspond to these positive eigenvalues.
| Genus | PCoord-1 | PCoord-2 | PCoord-3 |
|---|---|---|---|
| Acaste | 0.17 | -0.29 | -0.03 |
| Balizoma | 0.27 | -0.01 | -0.08 |
| Calymene | -0.18 | -0.22 | 0.10 |
| Ceraurus | 0.19 | -0.10 | -0.08 |
| Cheirurus | 0.04 | 0.28 | 0.37 |
| Cybantyx | -0.01 | 0.16 | -0.08 |
| Cybeloides | 0.14 | 0.00 | 0.02 |
| Dalmanites | 0.05 | -0.12 | -0.26 |
| Delphion | 0.17 | 0.23 | 0.22 |
| Ormathops | 0.27 | 0.07 | -0.17 |
| Phacopdina | 0.18 | 0.13 | -0.06 |
| Phacops | 0.11 | -0.05 | 0.32 |
| Placopoaria | -0.02 | -0.06 | -0.03 |
| Pricyclopyge | -0.07 | 0.45 | -0.12 |
| Ptychoparia | -0.30 | -0.52 | -0.25 |
| Rhenops | -0.26 | 0.28 | -0.48 |
| Sphaerexochus | 0.17 | -0.23 | 0.03 |
| Toxochasmops | -0.11 | -0.20 | 0.47 |
| Trimerus | -0.66 | 0.11 | 0.20 |
| Zacanthoides | -0.15 | 0.09 | -0.08 |
Unlike PCA in which the eigenvector values can be interpreted in terms of the geometry of the data, the eigenvectors of a PCoord analysis are simply scaling coefficients, useful only for representing the structure of the distance relations among taxa on the mutually orthogonal eigenvectors. This difference is underscored by the manner in which the PCoord scores are determined. Instead of using the eigenvector loadings to project the original data into the space defined by the new principal component axes (as is the case in PCA), the PCoord scores are determined simply by scaling the raw eigenvectors by the square root of the corresponding eigenvalues.
| Genus | PCoord-1 | PCoord-2 | PCoord-3 |
|---|---|---|---|
| Acaste | 1 | -3.41 | -0.21 |
| Balizoma | 23.07 | -0.15 | -0.97 |
| Calymene | -15.42 | -2.59 | 0.61 |
| Ceraurus | 16.24 | -1.22 | -0.52 |
| Cheirurus | 3.67 | 3.37 | 2.25 |
| Cybantyx | -1.26 | 1.92 | -0.48 |
| Cybeloides | 11.68 | -0.05 | 0.15 |
| Dalmanites | 0.00 | -1.48 | -1.61 |
| Delphion | 14.29 | 2.77 | 1.34 |
| Ormathops | 23.18 | 0.89 | -1.03 |
| Phacopdina | 15.05 | 1.53 | -0.38 |
| Phacops | 9.69 | -0.57 | 1.95 |
| Placopoaria | -1.79 | -0.75 | -0.17 |
| Pricyclopyge | -5.84 | 5.30 | -0.17 |
| Ptychoparia | -25.32 | -6.19 | -1.54 |
| Rhenops | -21.89 | 3.29 | -2.96 |
| Sphaerexochus | 14.46 | -2.69 | 0.18 |
| Toxochasmops | -9.58 | -2.35 | 2.86 |
| Trimerus | -56.36 | 1.27 | 1.23 |
| Zacanthoides | -12.65 | 1.12 | -0.47 |
In other words, while PCA uses its eigenvectors as a means to calculate its representation of the relations between objects or specimens, in PCoord analysis the eigenvectors are the representation of those relations. The rescaling operation is only needed because most eigenanalysis algorithms artificially adjust the lengths of the eigenvectors to a unit value. In a sense, this calculation simply restores the eigenvectors to their proper form.
Now, let us return to the question I asked at the beginning of the essay, how good are these scores at depicting structure of the original distance matrix. This can be assessed by calculating a new squared Euclidean Distance matrix based on the PCoord scores and comparing that to the original distance matrix. The first few entries for the distance matrix reproduced from scores on the first two PCoord axes is shown below.
| Acaste | Balizoma | Calymene | |
|---|---|---|---|
| Acaste | 0.000 | 78.027 | 917.634 |
| Balizoma | 78.027 | 0.000 | 1487.537 |
| Calymene | 917.634 | 1487.537 | 0.000 |
Comparison of this 'reproduced' distance matrix with Table 2 (above) shows that the first two PCoord axes provide a very good estimate of overall distance relations among specimens. A quantitative assessment of this 'fit' between the original and reproduced matrices can be gained by calculating the correlation between the two matrices. For this analysis that value is 0.9999. Finally, an image of these relations can be constructed by creating a scatterplot of scores on the first two PCoord axes (Fig. 1).
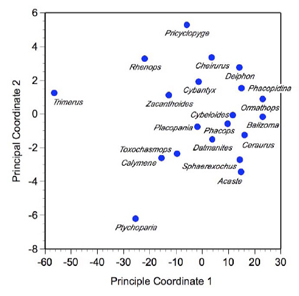
Figure 1. Scatterplot of the first two principal coordinates of the trilobite Euclidean Distance matrix.
If you've been following the column closely, this plot might look familiar. Indeed, it's almost identical in form to the plot we obtained for the space defined by the first two principal components of these data (see Fig. 7A, PalaeoMath 101, Principal Components Analysis (Eigenanalysis & Regression 5) from Newsletter 59). There are a few differences, though; some trivial and some important. If you go back and inspect Figure 7A of the PCA analysis essay you'll see that the relative positions of all taxa show on that plot are preserved by the PCoord results, but both axes have been reversed. This is a trivial difference. Remember, in eigenanalysis the direction of the axes is arbitrary, a result of the algorithm used to implement the eigenanalysis procedure, not the eigenanalysis concept itself. If we wished to change the direction of any PCoord or PCA axis we can simple multiply the scores along that axis by -1.0. It's perfectly 'legal' to do this and often facilitates the comparison of plots obtained from different datasets. The positions of the point clouds along the axes are also different for the PCoord and PCA results. This is another trivial difference. The axis system of the PCoord analysis is centered on the centroid of the trilobite data because the scaling step we undertook by applying Equation 7.4 to the raw squared Euclidean Distance matrix forced it to be so. Because we did not apply this transformation to the covariance matrix used as a basis for the PCA results, the PCA point cloud is located well away from the origin of the PCA coordinate system. Once again, bringing the coordinate systems into conformance would simply require that we mean-center the PCA scores, another operation that is eminently legal.
If we reversed the direction PCA axes and mean-centered the PCA scores for there trilobite taxa we'd notice another similarity, this one somewhat perplexing. We'd find that not only does the coordinate system change to a comparable position in both plots, but the scores of the trilobite taxa in the PCA space become identical with the scores of the these same taxa in the PCoord space. Even though we've now approached the analysis from two diametrically different points of view and used different matrices (embodying different definitions of 'relation'), we've obtained an identical result. What gives? If this is always the case, what's the point of a PCoord analysis?
To answer this question we need to go back to the beginning of the essay and return to first principles. As Gower (1966) showed"and as I mentioned above"a PCoord analysis based on the squared Euclidean Distance index is the dual of a covariance-based PCA analysis. These methods are inter-changeable and will always produce the same results in terms of the ordination of objects within the space of the new axes. This relation also extends to data that have been standardized prior to PCA or PCoord analysis. The point of a PCoord analysis, however is that whereas one is pretty much stuck with always performing a PCA analysis in this mode, PCoord analyses are much more flexible because they're not confined to being undertaken only using the squared Euclidean Distance metric.
For example, let's add a count of the number of pleural lobes on our trilobite specimens to the mean-centered trilobite data matrix.
| Genus | Body Length (mm) | Glabellar Length (mm) | Glabellar Width (mm) | Pleural Lobes |
|---|---|---|---|---|
| Acaste | -13.08 | -5.87 | -5.21 | 11 |
| Balizoma | -21.90 | -5.40 | -4.90 | 11 |
| Calymene | 15.47 | 1.54 | 1.74 | 13 |
| Ceraurus | -15.07 | -4.47 | -4.29 | 9 |
| Cheirurus | -4.48 | -0.04 | 3.13 | 12 |
| Cybantyx | 0.59 | 1.98 | 1.12 | 9 |
| Cybeloides | -11.09 | -2.98 | -2.17 | 12 |
| Dalmanites | -3.29 | -0.91 | -2.90 | 11 |
| Delphion | -14.41 | -2.45 | 0.03 | - |
| Ormathops | -22.34 | -4.34 | -4.46 | 9 |
| Phacopdina | -14.79 | -2.34 | 2.19 | 10 |
| Phacops | -8.89 | -4.07 | -0.79 | 11 |
| Placopoaria | 1.93 | 0.03 | -0.27 | 12 |
| Pricyclopyge | 3.89 | 5.61 | 4.00 | 5 |
| Ptychoparia | 25.95 | 2.88 | -0.27 | 12 |
| Rhenops | 19.72 | 9.63 | 4.12 | 10 |
| Sphaerexochus | -12.91 | -5.53 | -4.38 | 11 |
| Toxochasmops | 9.90 | -1.22 | 2.44 | 10 |
| Trimerus | 53.21 | 13.81 | 12.54 | 12 |
| Zacanthoides | 11.67 | 4.19 | 2.80 | 9 |
| Minimum | -22.34 | -5.87 | -5.21 | 5 |
| Maximum | 53.21 | 13.81 | 12.54 | 13 |
| Range | 75.55 | 19.68 | 17.75 | 8 |
| Mean | 0.00 | 0.00 | 0.00 | 10.47 |
| Variance | 346.89 | 27.33 | 18.27 | - |
Because this matrix mixes continuous variables and integer counts it could be argued that the Euclidean Distance index is not an appropriate means for estimating the among-specimen distance structure. The Gower index, however, is suited to Q-mode analysis of any data type or combination thereof.
Use of the Gower index results in a very different result from that of the previous analysis. First, the eigenvector decomposition of an appropriately scaled Gower matrix yields 18 positive eigenvalues, though note the step difference in terms of magnitude and percent contribution between eigenvalues 1-3 and 4-18.
| Principle Coord. | Eignevalue | Percent | Cum. Percent |
|---|---|---|---|
| 1 | 2.03 | 41.44 | 41.44 |
| 2 | 0.78 | 15.90 | 57.34 |
| 3 | 0.61 | 12.46 | 69.80 |
| 4 | 0.27 | 5.61 | 75.41 |
| 5 | 0.22 | 4.40 | 79.80 |
| 6 | 0.20 | 4.17 | 83.97 |
| 7 | 0.15 | 3.16 | 87.97 |
| 8 | 0.12 | 2.40 | 89.54 |
| 9 | 0.10 | 1.98 | 91.52 |
| 10 | 0.09 | 1.78 | 93.30 |
| 11 | 0.08 | 1.61 | 94.90 |
| 12 | 0.07 | 1.37 | 96.27 |
| 13 | 0.05 | 1.10 | 97.37 |
| 14 | 0.05 | 0.93 | 98.31 |
| 15 | 0.04 | 0.72 | 99.03 |
| 16 | 0.02 | 0.50 | 99.53 |
| 17 | 0.02 | 0.32 | 99.85 |
| 18 | 0.01 | 0.15 | 100.00 |
Analysis of the percent contribution of each principal coordinate to accounting for the association matrix suggests either a 3-axis or a 4-axis solution is appropriate. Subsequent calculation of the reproduced Gower matrix using scores on the first three PCoord axes results in a 92.4 percent replication of the observed structure. This means that the plots shown in Figure 2 represent a highly accurate picture of the distance-relations among our taxa on the basis of the four variables.
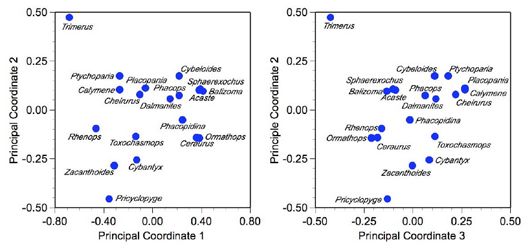
Figure 2. Scatterplot of the first three principal coordinates of the four-variable trilobite Gower matrix.
While it could be argued that these results are not strictly comparable to those of the three-variable analysis due to the absence of Delphion from this dataset and addition of the extra variable, inspection of Figure 1 suggests these changes did not have a strong influence on the overall character of the result. Compared to the previous ordination this result appears to confirm Trimerus and Pricyclopyge as outliers as well as exhibiting a comprehensive firming up of close correspondence (with respect to these variables) in several subsidiary groups (e.g., Acaste-Balizoma-Sphaerexochus, Ceraurus-Ormathops).
Whereas PCoord analysis can, at least in some forms, be considered an exact mirror or dual of PCA, Q-mode Factor Analysis (Q-FA) is definitely not the dual of classical or R-mode Factor Analysis. Once again, there are both trivial and an important reasons for this distinction. The trivial reason is that most textbooks recommend use of the Cosine θ association index as the best estimator of structural relations for Q-FA. Cosine θ is a safe choice as a basis for any Q-mode analysis in that it represents geometric relations between specimens in a way that is not sensitive to the magnitudes of the different variables used in their measurement-description and can even handle compositional data (data that have been artificially recalculated to sum to a constant value, such as proportions or percentages) effectively. The Cosine θ index is not, however, the only index that can be used in this context. Indeed, as with PCoord analysis, any distance or association index that produces a symmetric Q-mode matrix can be analyzed with the Q-FA procedure.
The important reason Q-FA is not the dual of R-FA is that, for all intents and purposes, Q-FA is nothing more than a PCoord analysis undertaken using the Cosine θ (or other) index. Recall my discussion of FA in the last newsletter. There I argued the only attribute truly unique to FA was the manner in which the scores of objects (e.g., specimens, localities) projected onto the FA axes were calculated. This complication is related to the statistical model on which FA is based and requires that only those aspects of variation that covary or are correlated with the FA model be used in the determination of the FA scores. In this way, the picture of relations among specimens obtained as a result of an FA analysis represents but an aspect of the variation exhibited by the data. In Q-FA none of this applies because"like PCoord"the scores are determined not as a result of a scaling of the original data, but as a result of the scaling of the eigenvector coefficients. Since no statistical model is used as the basis for a Q-FA analysis, the accepted Q-FA procedure is simply a routine variant of the procedure used in principle coordinates analysis; a PCoord analysis based on the Cosine θ association index.
This having been said, there is merit it taking a brief look at the results of such an analysis. Returning then to the original, three-variable trilobite data, the eigenanalysis decomposition of the Cosine θ matrix yields three eigenvalues, as follows.
| Principle Coord. | Eignevalue | Percent | Cum. Percent |
|---|---|---|---|
| 1 | 19.87 | 99.33 | 99.33 |
| 2 | 0.11 | 0.54 | 99.87 |
| 3 | 0.03 | 0.13 | 100.00 |
Note this is the most extreme decomposition we've seen yet, with all but 0.67 percent of the observed structural in formation being represented on a single axis. Such somewhat lopsided decompositions are often seen in PCoord results and you shouldn't be frightened by them. Just remember you're trying to understand the system of measurements, not use the mathematics to obtain a result that looks like someone else's results or (even worse) use numbers to paint a picture of your own pre-conceptions. In this instance the extremely large signal that's being pulled out by the first principal coordinate is due to the fact that (1) there are only three variables in the system, (2) all the variables are very highly associated with each other (see the fragment from the Cosine θ matrix, above), and (3) the Cosine θ index is essentially ignoring the component of inter-specific variation represented by size differences among the specimens, which is a very large component of this dataset.
Since there are only three coordinates (or 'factors', whichever you prefer) we'll take a look at all combinations in the plots (Fig. 3). There's no reason to calculate a reproduced Cosine θ matrix because everything there is to see about this result is shown below.
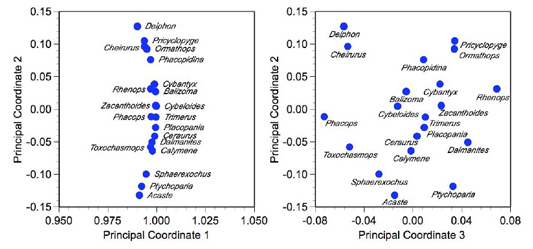
Figure 3. Scatterplot of the first three principal coordinates of the trilobite Cosine theta matrix.
The unusual character of the principal coordinate 1-2 plot is also often seen in Cosine θ-based PCoord analyses, especially for low-dimensional data. Somewhat counter-intuitively the data are arrayed along a well-defined arc centered on the coordinate (1,0). This arc represents a communality of 1.0 (see previous Factor Analysis essay from Newsletter 60). All points that fall on the arc have their positions determined by only two axes of the three-axis solution to this matrix. Those points plotting a bit behind the arc (e.g., Cheirurus, Phacops, Rhenops) exhibit a more complex pattern of variation that requires all three axes to place accurately. You can think of this arc as defining the surface of a clear, hollow sphere centered on the origin of the axis system. The plot of principal coordinates 1-2 is the image you'd see if you stood outside the sphere at a position tangent to its surface and at right angles to the mean data vector, while the principal coordinates 3-2 plot is the image you'd see if you stood in the centre of the coordinate system and looked out at the distribution of points on the surface of the sphere. Effectively, principal coordinate 1 is expressing that component of variation all the trilobites have in common whereas coordinates 2 and 3 express aspects of inter-sample difference.
Finally, note these plots are similar in some respects, but different in others to the previous representations of inter-specimen structural relations we've seen in this essay. Which representation is correct? The only reasonable answer is 'They all are!'. But that question is not specified in a very useful manner. The pertinent question is which of these indices best captures the distinctions between the trilobite specimens in which we are interested. Since this particular trilobite dataset exists only as means to illustrate mathematical techniques to a palaeontological audience, the 'which is correct' question has no answer in the context of the conversation we are having through these essays. Nevertheless, if you have data you wish to analyse by these methods, one presumes you collected the data for some purpose or to test some hypothesis. Selection of the appropriate Q-mode index will depend on the nature of the data you've collected and purpose(s) of the investigation. The decision will determine whether any PCoord result will be interesting or useful in the context of real data analysis situations. Everything else is just mathematical machinery.
Because principal coordinate analysis is not used as often as it should be, it's usually covered as an afterthought"if at all"in textbook treatments of data analysis methods, Add to this a genuine confusion among many authors as to what principal coordinates is and how it differs from other methods, as well as more-often-than-not, all but impenetrable mathematical presentations, and it's little wonder few palaeontologists understand, use, or even known about this method. The literature really is inordinately hard going. If principal coordinates analysis is not covered as a side issue to the presentation of PCA methods, it's usually discussed as an approach to multi-dimensional scaling (MDS). This is particularly unfortunate insofar as classical metric and non-metric MDS employs mathematical techniques that are wholly unrelated to those I've presented here, though in the end they try to accomplish something very similar. We'll take a look at MDS in an upcoming essay. Despite what you might read, though, MDS is very different from PCoord. Unfortunately, this general level of confusion is, if anything, elevated in the few statistical packages that implement PCoord analysis, or at least claim to. Of these, the only implementations I'd recommend are those included in NTSYS (http://www.exetersoftware.com/cat/ntsyspc/ntsyspc.html) and Syn-Tax (http://www.exetersoftware.com/cat/syntax/syntax.html). Likely there are other good principal coordinate programmes out there. If you know of any, let me know and I'll share the pertinent urls with the readership. I have PCoord routines I've written in Mathematica and would be happy to share with anyone who's interested. Drop me a line and let me know. Indeed, if there's sufficient interest within the PalAss community I might even be talked into coding and making available some simple applications for Windows and Mac platforms.
References
DAVIS, J. C. 2002. Statistics and data analysis in geology (third edition). John Wiley and Sons, New York, 638 pp.
GOWER, J. C. 1966. Some distance properties of latent roots and vectors used in multivariate analysis. Biometrika, 53, 588-589.
JACKSON, J. E. 1991. A user's guide to principal components. John Wiley & Sons, New York, 592 pp.
JOLLIFFE, I. T. 2002. Principal component analysis (second edition). Springer-Verlag. New York, 516 pp.
MANLEY, B. F. J. 1994. Multivariate statistical methods: a primer. Chapman & Hall, Bury, St. Edmonds, Suffolk, 215 pp.
PEILOU, E. C. 1984. The interpretation of ecological data. John Wiley & Sons, New York, 263 pp.
REYMENT, R. A. and JÖRESKOG, K. G. 1993. Applied factor analysis in the natural sciences. Cambridge University Press, Cambridge, 371 pp
Footnotes
1These values were calculated using the variable ranges for the entire trilobite dataset (see Table 1). See the PalaeoMath 101 Spreadsheet, PCoord Worksheet for full details.
2Usually application of Equation 7.4 reduces the number of positive eigenvalues to one less than the number of variables measured. That hasn’t happened in this analysis due to the small number of variables measured, coupled with rounding error. Note the relative contribution of the third principal coordinate is very small.
3Delphion measurements were taken on a specimen that only consisted of a cranidium. Accordingly, this taxon will be eliminated from the four-variable analysis.
